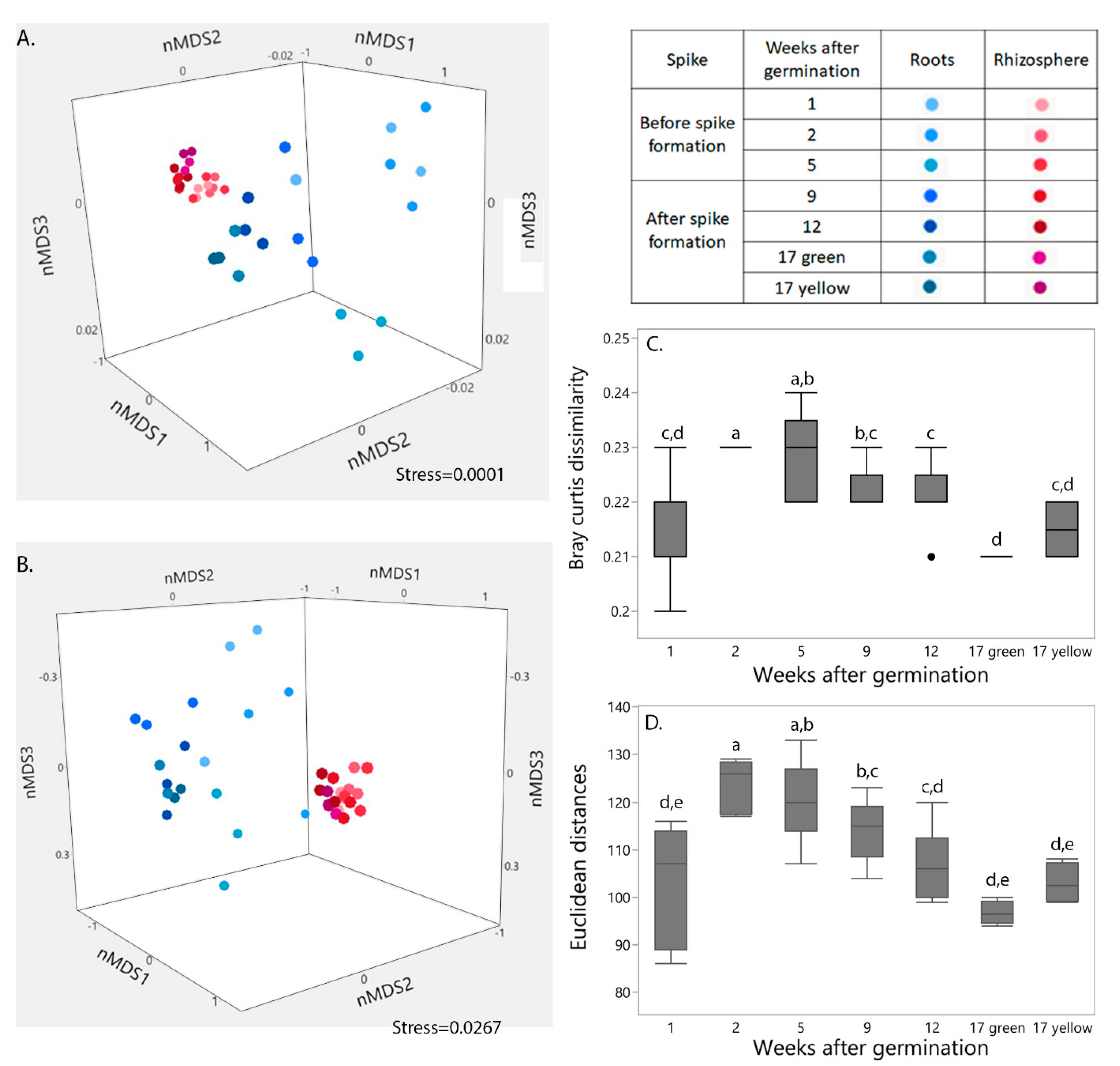
IJMS | Free Full-Text | Spike Formation Is a Turning Point Determining Wheat Root Microbiome Abundance, Structures and Functions

V8 BOX | New Alternative for VphoneGaga Vmos Pro X8 SandBox F1VM | V8 Box ROOT PlayStore 32/64bit - YouTube
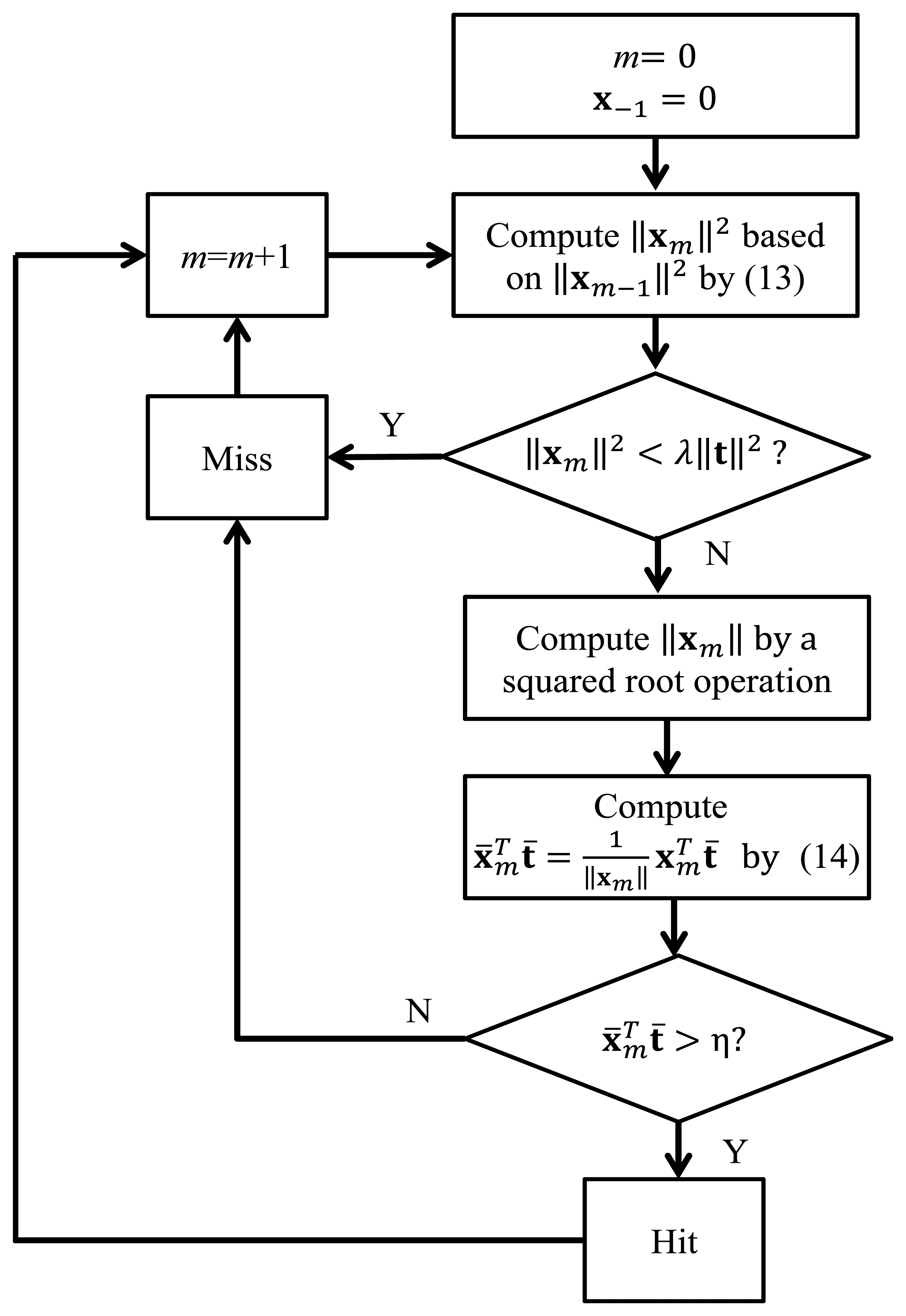
Sensors | Free Full-Text | Spike Detection Based on Normalized Correlation with Automatic Template Generation

Systematic virtual screening in search of SARS CoV-2 inhibitors against spike glycoprotein: pharmacophore screening, molecular docking, ADMET analysis and MD simulations | SpringerLink
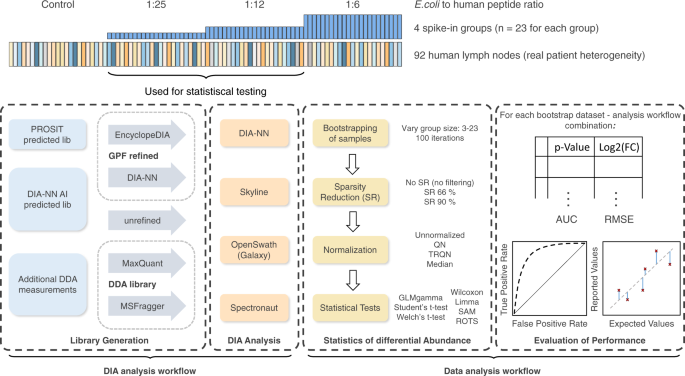
Benchmarking of analysis strategies for data-independent acquisition proteomics using a large-scale dataset comprising inter-patient heterogeneity | Nature Communications